2.4: What is the Usefulness of an Ontology?
- Page ID
- 6403
Now that we have some idea of ontologies, let us have a look at where they are being used. Ontologies for information systems were first proposed to contribute to solving the issues with data integration: an ontology provides the common vocabulary for the applications that is at one level of abstraction higher up than conceptual data models such as EER diagrams and UML Class Diagrams. Over the years, it has been used also for other purposes. We start with two distinct scenarios of data integration where ontologies play a central role, and subsequently describe other scenarios where ontologies are an important part of the solution.
Data and Information System Integration
Figure 1.3.1 sketches the idea of the ontology-driven schema-based data integration and Figure 1.3.2 further below shows an example of data-based data integration that we shall elaborate on in the next two subsections.
.png?revision=1&size=bestfit&width=1104&height=636)
Figure 1.3.1: Sketch of an ontology-based application integration scenario. Bottom: different implementations, such as relational databases and OO software; Centre: conceptual data models tailored to the application (a section of an EER, ORM, and UML diagram, respectively); Top: an ontology that provides a shared common vocabulary for interoperability among the applications. See text for explanation.
Integrating Legacy Systems
In the setting of ontology-driven schema-based (and conceptual data model-based) data integration, a typical situation is as follows. You have several databases containing data on the same topic. For instance, two universities join forces into one: each university had its own database with information about students, yet, as the new mega-university, there has to be one single database to manage the data of all students. This means that the two databases have to be integrated somehow. A similar situation occurs oftentimes in industry, especially due to mergers and acquisitions, in government due to the drive for e-Government services to the citizens of the country, or attempting to develop a software system for integrated service delivery, as well as in healthcare due to a drive for electronic health records that need to combine various systems, say, a laboratory database with the doctor’s database, among other scenarios.
While the topic of data integration deserves its own textbook7, we focus here only on the ontology-driven aspect. Let us assume we have the relational databases and therewith at least their respective physical schemas, and possibly also the relational model and even the respective conceptual models, and also some objectoriented application software on top of the relational database. Their corresponding conceptual data models are tailored to the RDBMS/OO application and may or may not be modelled in the same conceptual data modelling language; e.g., one could be in EER, another in ORM, in UML and so forth. The example in Figure 1.3.1 is a sketch of such a situation about information systems of flower shops, where at the bottom of the figure we have two databases and one application that has been coded in C++. In the layer above that, there is a section of their respective conceptual data models: we have one in EER with bubble-notation, one in ORM, and one UML Class Diagram. Each conceptual data model has “Flower” and “Colour” included in some way: in the UML Class diagram, the colour is an attribute of the flower, i.e., Color → FlowerXString (that actually uses only the values of the Pantone System) and similarly in the EER diagram (but then without the data type), and in the ORM diagram the colour is a value type (unary predicate) Kleur with an additional relation to the associated datatype colour region in the spectrum with as data type real. Clearly, the notion of the flower and its colour is the same throughout, even though it is represented differently in the conceptual data models and in the implementations. It is here that the ontology comes into play, for it is the place to assert exactly that underlying, agreed-upon notion. It enables one to assert that:
- EER’s and UML diagram’s
Flowerand ORM’sBloem‘means’ Flower in the domain ontology8, which is indicated with the red dashed arrows. - EER’s
Colour, ORM’sKleurand UML’sColordenote the same kind of thing, albeit at one time it is represented as a unary predicate (in ORM) and other times it is a binary relation with a data type, i.e., an attribute. Their ‘mappings’ to the entity in the ontology (green dashed arrows), Colour, indicates that agreement. - There is no agreement among the conceptual models when it comes to the data type used in the application, yet they may be mapped into their respective notion in an ontology (purple dashed arrows). For instance, the ColourRegion for the values of the colour(s) in the colour spectrum is a PhysicalRegion, and one might say that the PantoneSystem of colour encoding is an AbstractRegion.
The figure does not include names of relationships in the conceptual data model, but they obviously can be named at will; e.g., heeftKleur (‘hasColour’) in the ORM diagram. Either way, there is, from an ontological perspective, a specific type of relation between the class and its attribute: one of dependency or inherence, i.e., that specific colour instance depends on the existence of the flower, for if that particular flower does not exist, then that specific instance of colour does not exist either. An ontology can provide those generic relations, too. In the sketch, this happens to be the \(qt\) relationship between enduring objects (like flowers) and the qualities they have (like their colour), and from the quality to the value regions (more precisely: qualia), the relation is called \(ql\) in Figure 1.3.1.
Although having established such links does not complete the data integration, it is the crucial step—the rest has become, by now, largely an engineering exercise.
.png?revision=1&size=bestfit&width=945&height=696)
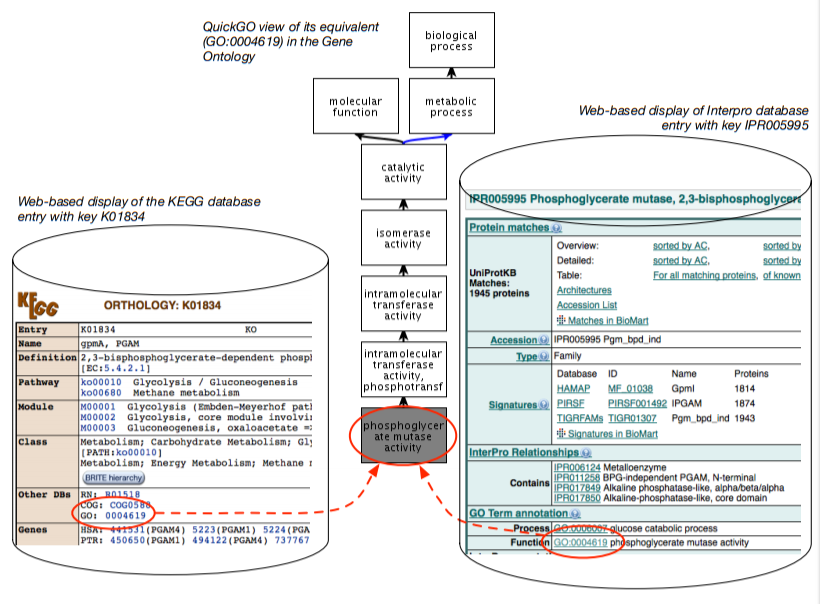
Figure 1.3.2: Illustration of ontology-based data-level integration: two databases, the KEGG and InterPro, with a web-based front-end, and each database has its data (each tuple in the database, where possible) annotated with a term from the Gene Ontology.
Data-Level Data Integration
While in computer science the aforementioned approach to data integration was under investigation, domain experts in molecular biology needed a quick and practical solution to the data integration problem, urgently. Having noticed the idea of ontologies, they came up with another approach, being interoperability at the instance-level, tuple-by-tuple, or even cell-by-cell, and that with multiple databases over the Internet instead of the typical scenario of RDBMSs within an organisation. This can be achieved with lightweight ontologies, or structured controlled vocabularies.
The basic idea is illustrated in Figure 1.3.2. There are multiple databases, which in the figure are the KEGG and InterPro databases. In the KEGG database, there is a tuple with as key K01834 and it has several attributes (columns in the table in the relational database), such as the name (gpmA), and further down in the display there is an attribute Other DBs, which has as entry GO:0004619; i.e., there is a tuple in the table along the line of <K01834, ..., GO:0004619>. In the InterPro database, we have a similar story but then for the entity with the key IPR005995, where there is a section “GO Term Annotation” with an attribute function that has GO:0004619; i.e., a tuple <IPR005995, ..., GO:0004619>. That is, they are clearly distinct tuples—each with their separate identifier from a different identifier scheme, with different attributes, one physically stored in a database in Japan and the other in the USA—yet they actually talk about the same thing: GO:0004619, which is the identifier for Phosphoglycerate Mutase Activity.
The “GO:0004619” is an identifier for a third artefact: a class in the Gene Ontology (GO) [Gen00]. The GO is a structured controlled vocabulary that contains the concepts and their relationship that the domain experts agree upon to annotate genes with; the GO contains over 40000 concepts by now. The curators of the two databases each annotated their entity with a term from the GO, and thereby they assert they have to do with that same thing, and therewith have created an entitylevel linking and interoperability through the GO. Practically, on top of that, these fields are hyperlinked (in the soft copy: blue text in KEGG and green underlined text in the screenshot of the InterPro entry), so that a vast network of data-level interlinked databases has been created. Also, the GO term is hyperlinked to the GO file online, and in this way, you can browse from database to database availing of the terms in the ontology without actually realising they are wholly different databases. Instead, they appear like one vast network of knowledge.
There are many more such ontologies, and several thousand databases that are connected in this way, not only thanks to ontologies, but where the ontology serves as the essential ingredient to the data integration. Some scientific journals require the authors to use those terms from the ontologies when they write about their discoveries, so that one more easily can find papers about the same entity9.
Trying to Prevent Interoperability Problems
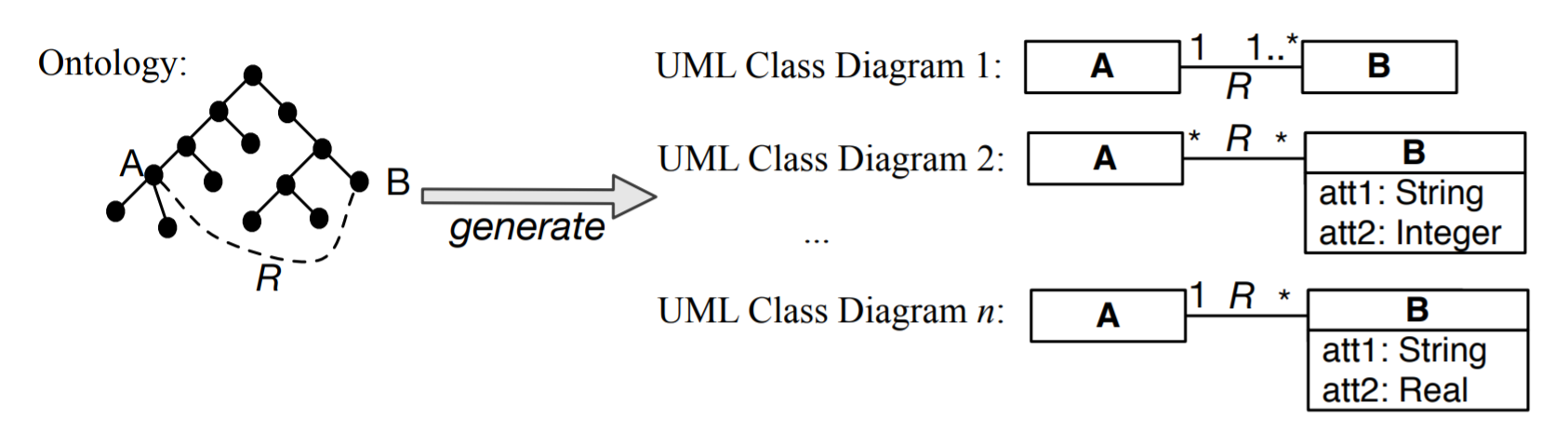
A related topic in the data integration scenarios, is trying to prevent the integration problems form happening in the first place. This may be done though generating conceptual models for related new applications based on the knowledge represented in the ontology [EGOMA06, JDM03, SS06]. This is a bit alike the Enterprise Models you may have come across in information system design. In this way, interoperability is guaranteed upfront because the elements in the new conceptual data models are already shared thanks to the link with the same ontology. For instance, some relation R between A and B, as is depicted in Figure 1.3.3 (e.g., an enrols relation between Student and Course), is reused across the conceptual data models, yet each model may have its own additional constraints and data types for attributes. For instance, in one university (hence, student information management system), students may not register for more than six courses and have to be registered for at least one to count as student, whereas at another university, a student may well decide not to be registered for any course at a particular time during the year and still count as a registered student. Whichever rules there may be at each individual university, the systems do agree on the notions of Student, Course, and enrols. Thus, the ontology provides the shared common vocabulary for interoperability among the applications.
.png?revision=1)
Figure 1.3.3: Basic illustration of taking information from an ontology and using it for several conceptual data models (here: UML Class Diagrams), where the constraints may be refined or attributes added, yet sharing the semantics of A, B, and R.
Ontologies as Part of a Solution to Other Problems
Over the years, ontologies have been shown to be useful in a myriad of other application scenarios; among others, negotiation between software services, mediation between software agents, bringing more quality criteria into conceptual data modelling to develop a better model (hence, a better quality software system), orchestrating the components in semantic scientific workflows, e-learning, ontology-based data access, information retrieval, management of digital libraries, improving the accuracy of question answering systems, and annotation and analysis of electronic health records, to name but a few. Four of them are briefly illustrated in this section.
e-Learning
The ‘old-fashioned’ way of e-learning is a so-called content-push: the lecturer sends out softcopies of the notes, slides, answers to the solutions, and perhaps the video recordings of the lectures, and the student consumes it. This is a one-size-fits-all approach regardless the student’s background with acquired knowledge and skills, and learning preferences and habits, which cannot be assumed to be homogeneous in an e-learning setting, or at least much less so than with respect to your fellow students in the ontology engineering class. A more sophisticated way for e-learning is adaptive e-learning, which tailors the contents to the student based on prior knowledge and learning habits. To be able to automatically tailor the offering to the student, one has to develop a ‘smart’ e-learning application that can figure out what kind of student is enrolled. Put differently: students have certain properties (part-time/full-time student, age, undergraduate degree, etc.), the learning objects have to be annotated (by skill level and topic), and user logs have to be categorised according to type of learning pattern, and based on that the material and presentation can be adjusted, like skipping the section on first order logic and delve deeper into, or spend more time on, ontology engineering and modelling if you have a mathematics background, whereas a philosopher may crave for more content about foundational ontologies but skip reverse engineering of relational databases, or offer a student more exercises on a topic s/he had difficulties with. This requires knowledge representation—of the study material, questions, answers, students’ attributes, learning approaches—and automated reasoning to classify usage pattern and student, and annotated content, i.e., using ontologies and knowledge bases to make it work solidly and in a repeatable way. See, e.g., [HDN04] as a start for more details on this topic.
Deep Question Answering with Watson
Watson10 is a sophisticated question answering engine that finds answers to trivia/general knowledge questions for the Jeopardy! TV quiz that, in the end, did consistently outperform the human experts of the game. For instance, a question could be “who is the president of South Africa?”: we need algorithms to parse the question, such as that ‘who’ indicates the answer has to be a person and a named entity, it needs to be capable to detect that South Africa is a country and what a country is, and so on, and then have some kind of a look-up service in knowledge bases and/or natural language documents to somehow find the answer by relating ‘president’, ‘South Africa’ and ‘Cyril Ramaphosa’ and that he is the current president of the country. An ontology can then be used in the algorithms of both the understanding of the question and finding the right answer11 and integrating data sources and knowledge, alongside natural language processing, statistical analysis and so on, comprising more than 100 different techniques12. Thus, a key aspect of the system’s development was that one cannot go in a linear fashion from natural language to knowledge management, but have to use an integration of various technologies, including ontologies, to make a successful tool.
Digital humanities
Some historians and anthropologists in the humanities try to investigate what happened in the Mediterranean basin some 2000 years ago, aiming to understand food distribution systems. Food was stored in pots (more precisely: an amphora) that had engravings on it with text about who, what, where etc. and a lot of that has been investigated, documented, and stored in multiple resources, such as in databases. None of the resources cover all data points, but to advance research and understanding about it and food trading systems in general, it has to be combined and made easily accessible to the domain experts. That is, essentially it is an instance of a data access and integration problem. Also, humanities researchers are not at all familiar with writing SQL queries, so that would need to be resolved as well, and in a flexible way so that they would not have to be dependent on the availability of a system administrator.
A recent approach, of which the technologies have been maturing, is OntologyBased Data Access (OBDA). The general idea of OBDA applied to the Roman Empire Food system is shown in Figure 1.3.4. There are the data sources, which are federated (one ‘middle layer’, though still at the implementation level). The federated interface has mapping assertions to elements in the ontology. The user then can use the terms of the ontology (classes and their relations and attributes) to query the data, without having to know about how the data is stored and without having to write page-long SQL queries. For instance, a query “retrieve inscriptions on amphorae found in the city of ‘Mainz’ containing the text ‘PNN”” would use just the terms in the ontology, say, Inscription, Amphora, City, found in, and inscribed on, and any value constraint added (like the PNN), and the OBDA system takes care of the rest to return the answer.
Semantic Scientific Workflows
Due to the increase in a variety of equipment, their speed, and decreasing price, scientists are generating more data than ever, and are collaborating more. This data has to be analysed and managed. In the early days, and, to some extent, to this day, many one-off little tools were developed, or simply scripted together with PERL, Ruby on Rails, or Python, used once or a few times and then left for what it was. This greatly hampers repeatability of experiments, insight in the provenance of the data, and does not quite follow a methodological approach for so-called in silico biology research. Over the past 10 years, comprehensive software and hardware infrastructures have been, and are being, built to fix these and related problems. Those IT ‘workbenches’ (as opposed to the physical ones in the labs) are realised in semantic scientific workflow systems. An example of a virtual bench is Taverna [GWG+07], which in the meantime has gone ‘mainstream’ as a project within the Apache Software Foundation13 [WHF+13]. This, in turn can be extended further; e.g., to incorporate data mining in the workflow to go with the times of Big Data, as depicted in Figure 1.3.5. This contains ontologies of both the subject domain where it is used as well as ontologies about data mining itself that serve to find appropriate models, algorithms, datasets, and tools for the task at hand, depicted in Figure 1.3.6. Thus, we have a large software system—the virtual workbench—to facilitate scientists to do their work, and some of the components are ontologies for integration and data analysis across the pipeline.
.png?revision=1)
Figure 1.3.4: OBDA in the EPnet system (Source: [CLM+16])
Now that we have seen some diverse examples, this does not mean that ontologies are the panacea for everything, and some ontologies are better suitable to solve one or some of the problems, but not others. Put differently, it is prudent to keep one’s approach to engineering: conduct a problem analysis first, collect the requirements and goals, and then assess if an ontology indeed is part of the solution or not. If it is part of the solution, then we enter in the area of ontology engineering.
Success Stories
To be able to talk about successes of ontologies, and its incarnation with Semantic Web Technologies in particular, one first needs to establish when something can be deemed a success, when it is a challenge, and when it is an outright failure. Such measures can be devised in an absolute sense—compare technology \(x\) with an ontology-mediated one: does it outperform on measure \(y\)?—and relative—to whom is technology \(x\) deemed successful?
A major success story of the development and use of ontologies for data linking and integration is the Gene Ontology [Gen00], its offspring, and subsequent coordinated evolution of ontologies [SAR+07]. These frontrunners from the Gene Ontology Consortium14 and their colleagues in bioinformatics were adopters of some of the Semantic Web ideas even before Berners-Lee, Hendler, and Lassila wrote their Scientific American paper in 2001 [BLHL01], even though they did not formulate their needs and intentions in the same terminology: they did want to have shared, controlled vocabularies with the same syntax to facilitate data integration—or at least interoperability—across Web-accessible databases, have a common space for identifiers, it needing to be a dynamic, changing system, to organize and query incomplete biological knowledge, and, albeit not stated explicitly, it all still needed to be highly scalable [Gen00]. The results exceeded anyone’s expectations in its success for a range of reasons. Many tools for the Gene Ontology (GO) and its common Knowledge Representation format, .obo, have been developed, and other research groups adopted the approach to develop controlled vocabularies either by extending the GO, e.g., rice traits, or adding their own subject domain, such as zebrafish anatomy and mouse developmental stages. This proliferation, as well as the OWL development and standardisation process that was going on at about the same time, pushed the goal posts further: new expectations were put on the GO and its siblings and on their tools, and the proliferation had become a bit too wieldy to keep a good overview what was going on and how those ontologies would be put together. Put differently, some people noticed the inferencing possibilities that can be obtained from moving from a representation in obo to one in OWL and others thought that some coordination among all those obo bio-ontologies would be advantageous given that post-hoc integration of ontologies of related and overlapping subject domains is not easy. Thus came into being the OBO Foundry to solve such issues, proposing an approach for coordinated evolution of ontologies to support biomedical data integration [SAR+07] within the OBO Foundry Project15.
People in related disciplines, such as ecology, have taken on board experiences of these very early adopters, and instead decided to jump on board after the OWL standardization. They, however, were not only motivated by data(base) integration. Referring to Madin et al’s paper [MBSJ08], I highlight three points they made:
– “terminological ambiguity slows scientific progress, leads to redundant research efforts, and ultimately impedes advances towards a unified foundation for ecological science”, i.e., identification of some serious problems they have in ecological research;
– “Formal ontologies provide a mechanism to address the drawbacks of terminological ambiguity in ecology”, i.e., what they expect that ontologies will solve for them (disambiguation); and
– “fill an important gap in the management of ecological data by facilitating powerful data discovery based on rigorously defined, scientifically meaningful terms”, i.e., for what purpose they want to use ontologies and any associated computation (discovery using automated reasoning).
That is, ontologies not as a—one of many possible—‘tool’ in the engineering infrastructure, but as a required part of a method in the scientific investigation that aims to discover new information and knowledge about nature (i.e., in answering the who, what, where, when, and how things are the way they are in nature). Success in inferring novel biological knowledge has been achieved with classification of protein phosphatases [WSH07], precisely thanks to the expressive ontology and its automated reasoning services.
Good quality ontologies have to built and maintained, though, and there needs to be working infrastructure for it. This textbook hopefully will help you with that.
Footnotes
7the ‘principles of...’ series may be a good start; e.g., [DHI12].
8that in this case is linked to a foundational ontology, DOLCE, and there it is a subclass of a Non-Agentive Physical Object; we return to this in Block II
9It used to be a sport among geneticists to come up with cool names for the genes they discovered (e.g., “Sonic hedgehog”); when a gene was independently recovered, each research team typically had given the gene a different name, which can end up as a Tower of Babel of its own that hampered progress in science. The GO and similar ontologies resolve that issue.
10http://en.Wikipedia.org/wiki/Watson_(computer)
11On a much more modest scale, as well as easier accessible and shorter to read, Vila and Ferrández describe this principle and demonstrated benefits for their Spanish language based question-answering system in the agricultural domain [VF09].
12ftp://public.dhe.ibm.com/common/ssi/...W03061USEN.PDF
13https://taverna.incubator.apache.org/


